countData <- read.csv("GSE37704_featurecounts.csv", row.names = 1)
colData <- read.csv("GSE37704_metadata.csv")Class 14: RNASeq Mini-Project
Background
The Data for today’s mini-project comes from a knock-down study of an important HoX gene.
Data Import
Clean up (data tidying)
countData <- as.matrix(countData[,-1])
head(colData) id condition
1 SRR493366 control_sirna
2 SRR493367 control_sirna
3 SRR493368 control_sirna
4 SRR493369 hoxa1_kd
5 SRR493370 hoxa1_kd
6 SRR493371 hoxa1_kdhead(countData) SRR493366 SRR493367 SRR493368 SRR493369 SRR493370 SRR493371
ENSG00000186092 0 0 0 0 0 0
ENSG00000279928 0 0 0 0 0 0
ENSG00000279457 23 28 29 29 28 46
ENSG00000278566 0 0 0 0 0 0
ENSG00000273547 0 0 0 0 0 0
ENSG00000187634 124 123 205 207 212 258countData <- countData[rowSums(countData) != 0, ]
head(countData) SRR493366 SRR493367 SRR493368 SRR493369 SRR493370 SRR493371
ENSG00000279457 23 28 29 29 28 46
ENSG00000187634 124 123 205 207 212 258
ENSG00000188976 1637 1831 2383 1226 1326 1504
ENSG00000187961 120 153 180 236 255 357
ENSG00000187583 24 48 65 44 48 64
ENSG00000187642 4 9 16 14 16 16DESeq Analysis
Setting up the DESeq object
library(DESeq2)Running Deseq
dds = DESeqDataSetFromMatrix(countData=countData,
colData=colData,
design=~condition)Warning in DESeqDataSet(se, design = design, ignoreRank): some variables in
design formula are characters, converting to factorsGetting results
dds = DESeq(dds)estimating size factorsestimating dispersionsgene-wise dispersion estimatesmean-dispersion relationshipfinal dispersion estimatesfitting model and testingddsclass: DESeqDataSet
dim: 15975 6
metadata(1): version
assays(4): counts mu H cooks
rownames(15975): ENSG00000279457 ENSG00000187634 ... ENSG00000276345
ENSG00000271254
rowData names(22): baseMean baseVar ... deviance maxCooks
colnames(6): SRR493366 SRR493367 ... SRR493370 SRR493371
colData names(3): id condition sizeFactorres = results(dds)summary(res)
out of 15975 with nonzero total read count
adjusted p-value < 0.1
LFC > 0 (up) : 4349, 27%
LFC < 0 (down) : 4396, 28%
outliers [1] : 0, 0%
low counts [2] : 1237, 7.7%
(mean count < 0)
[1] see 'cooksCutoff' argument of ?results
[2] see 'independentFiltering' argument of ?resultsVolcano Plot
library(ggplot2)
ggplot(res) +
aes(log2FoldChange,
-log(padj)) +
geom_point()Warning: Removed 1237 rows containing missing values or values outside the scale range
(`geom_point()`).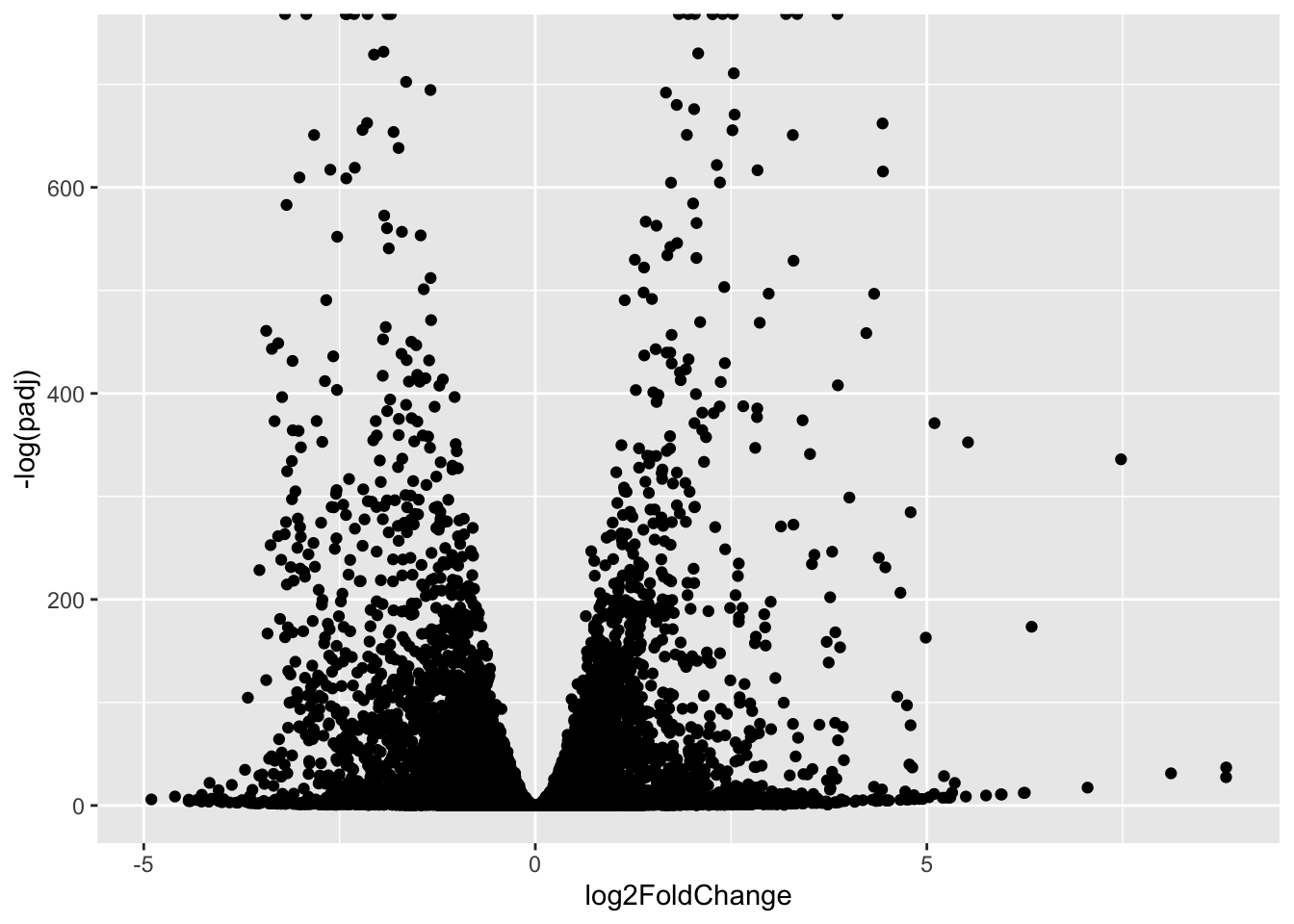
mycols <- rep("lightblue", nrow(res) )
mycols[ abs(res$log2FoldChange) > 2 ] <- "green"
mycols[ res$padj > 0.01 ] <- "lightblue"
ggplot(res) +
aes(log2FoldChange,
-log(padj)) +
geom_point(col= mycols) +
xlab("Log2(FoldChange)") +
ylab("-Log(P-value)") +
geom_vline(xintercept = c(-2,2)) +
geom_hline(yintercept = -log10(0.01))Warning: Removed 1237 rows containing missing values or values outside the scale range
(`geom_point()`).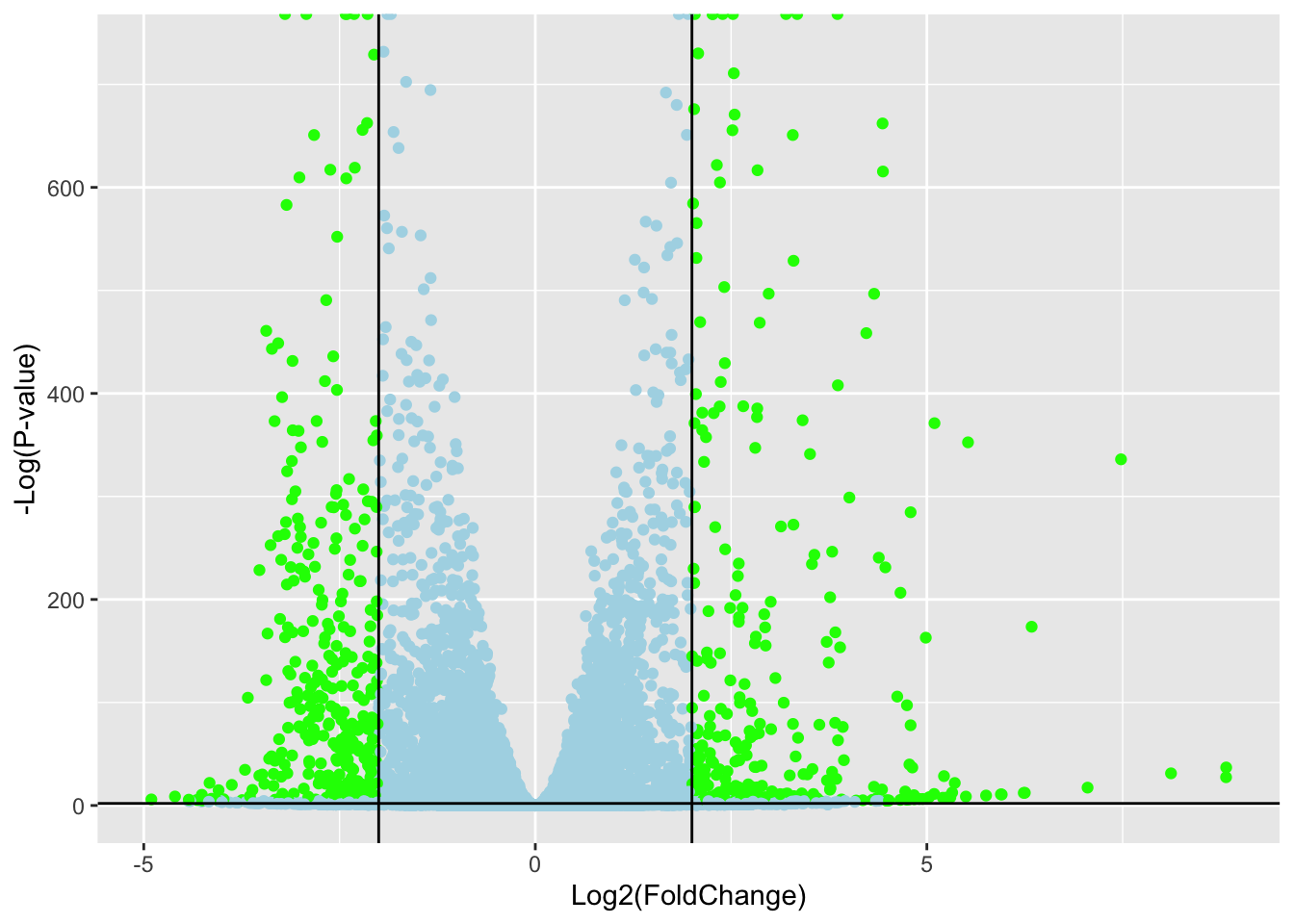
Add Annotation
library("AnnotationDbi")
library("org.Hs.eg.db")columns(org.Hs.eg.db) [1] "ACCNUM" "ALIAS" "ENSEMBL" "ENSEMBLPROT" "ENSEMBLTRANS"
[6] "ENTREZID" "ENZYME" "EVIDENCE" "EVIDENCEALL" "GENENAME"
[11] "GENETYPE" "GO" "GOALL" "IPI" "MAP"
[16] "OMIM" "ONTOLOGY" "ONTOLOGYALL" "PATH" "PFAM"
[21] "PMID" "PROSITE" "REFSEQ" "SYMBOL" "UCSCKG"
[26] "UNIPROT" res$symbol <- mapIds(org.Hs.eg.db,
keys=rownames(res),
keytype="ENSEMBL",
column="SYMBOL",
multiVals="first")'select()' returned 1:many mapping between keys and columnsres$entrez <- mapIds(org.Hs.eg.db,
keys=rownames(res),
keytype="ENSEMBL",
column="ENTREZID",
multiVals="first")'select()' returned 1:many mapping between keys and columnsres$name <- mapIds(org.Hs.eg.db,
keys=row.names(res),
keytype="ENSEMBL",
column="GENENAME",
multiVals="first")'select()' returned 1:many mapping between keys and columnshead(res, 10)log2 fold change (MLE): condition hoxa1 kd vs control sirna
Wald test p-value: condition hoxa1 kd vs control sirna
DataFrame with 10 rows and 9 columns
baseMean log2FoldChange lfcSE stat pvalue
<numeric> <numeric> <numeric> <numeric> <numeric>
ENSG00000279457 29.913579 0.1792571 0.3248216 0.551863 5.81042e-01
ENSG00000187634 183.229650 0.4264571 0.1402658 3.040350 2.36304e-03
ENSG00000188976 1651.188076 -0.6927205 0.0548465 -12.630158 1.43990e-36
ENSG00000187961 209.637938 0.7297556 0.1318599 5.534326 3.12428e-08
ENSG00000187583 47.255123 0.0405765 0.2718928 0.149237 8.81366e-01
ENSG00000187642 11.979750 0.5428105 0.5215598 1.040744 2.97994e-01
ENSG00000188290 108.922128 2.0570638 0.1969053 10.446970 1.51282e-25
ENSG00000187608 350.716868 0.2573837 0.1027266 2.505522 1.22271e-02
ENSG00000188157 9128.439422 0.3899088 0.0467163 8.346304 7.04321e-17
ENSG00000237330 0.158192 0.7859552 4.0804729 0.192614 8.47261e-01
padj symbol entrez name
<numeric> <character> <character> <character>
ENSG00000279457 6.86555e-01 NA NA NA
ENSG00000187634 5.15718e-03 SAMD11 148398 sterile alpha motif ..
ENSG00000188976 1.76549e-35 NOC2L 26155 NOC2 like nucleolar ..
ENSG00000187961 1.13413e-07 KLHL17 339451 kelch like family me..
ENSG00000187583 9.19031e-01 PLEKHN1 84069 pleckstrin homology ..
ENSG00000187642 4.03379e-01 PERM1 84808 PPARGC1 and ESRR ind..
ENSG00000188290 1.30538e-24 HES4 57801 hes family bHLH tran..
ENSG00000187608 2.37452e-02 ISG15 9636 ISG15 ubiquitin like..
ENSG00000188157 4.21963e-16 AGRN 375790 agrin
ENSG00000237330 NA RNF223 401934 ring finger protein ..res = res[order(res$pvalue),]
write.csv(res, file="deseq_results.csv")##Pathway Analysis
library(pathview)
library(gage)
library(gageData)
data(kegg.sets.hs)
data(sigmet.idx.hs)
foldchanges = res$log2FoldChange
names(foldchanges) = res$entrez
head(foldchanges) 1266 54855 1465 2034 2150 6659
-2.422719 3.201955 -2.313738 -1.888019 3.344508 2.392288 head(kegg.sets.hs,5)$`hsa00232 Caffeine metabolism`
[1] "10" "1544" "1548" "1549" "1553" "7498" "9"
$`hsa00983 Drug metabolism - other enzymes`
[1] "10" "1066" "10720" "10941" "151531" "1548" "1549" "1551"
[9] "1553" "1576" "1577" "1806" "1807" "1890" "221223" "2990"
[17] "3251" "3614" "3615" "3704" "51733" "54490" "54575" "54576"
[25] "54577" "54578" "54579" "54600" "54657" "54658" "54659" "54963"
[33] "574537" "64816" "7083" "7084" "7172" "7363" "7364" "7365"
[41] "7366" "7367" "7371" "7372" "7378" "7498" "79799" "83549"
[49] "8824" "8833" "9" "978"
$`hsa01100 Metabolic pathways`
[1] "10" "100" "10007" "100137049" "10020" "10026"
[7] "100510686" "10063" "10157" "10170" "10195" "10201"
[13] "10229" "10312" "10317" "10327" "10331" "1036"
[19] "10380" "10390" "1040" "10400" "10402" "10423"
[25] "10449" "10476" "10554" "10555" "10558" "1056"
[31] "10588" "10606" "10621" "10622" "10623" "10632"
[37] "10654" "1066" "10678" "10682" "10690" "10714"
[43] "10720" "10768" "10797" "10826" "10841" "10855"
[49] "10873" "10901" "10905" "10941" "10975" "10993"
[55] "10998" "11019" "11041" "1109" "11112" "11128"
[61] "1119" "1120" "11226" "11227" "11232" "112483"
[67] "11253" "11282" "11285" "113026" "11320" "11343"
[73] "113451" "113612" "114805" "1152" "1158" "1159"
[79] "1160" "116285" "117248" "119548" "120227" "121278"
[85] "122481" "122622" "123099" "123745" "123876" "124"
[91] "124454" "124975" "125" "125061" "125965" "125981"
[97] "126" "126328" "126792" "127" "127124" "128"
[103] "128869" "129607" "129642" "130" "130013" "131"
[109] "1312" "131669" "132" "132158" "1327" "132789"
[115] "1329" "1337" "1339" "1340" "134147" "1345"
[121] "1349" "1350" "1351" "135152" "1352" "1353"
[127] "1355" "1371" "1373" "137964" "138050" "138429"
[133] "139596" "140838" "1431" "144193" "144245" "145226"
[139] "146664" "1491" "15" "1503" "150763" "151056"
[145] "151531" "1537" "154141" "1543" "1544" "1548"
[151] "1549" "155066" "1551" "1553" "1555" "1557"
[157] "1558" "1559" "1562" "1571" "1573" "157506"
[163] "1576" "1577" "1579" "158" "1581" "1582"
[169] "1583" "1584" "1585" "1586" "1588" "1589"
[175] "159" "1593" "1594" "1595" "160287" "1603"
[181] "1606" "1607" "1608" "160851" "1609" "1610"
[187] "1621" "162417" "162466" "1629" "1633" "1635"
[193] "1638" "1644" "1650" "166929" "168391" "169355"
[199] "170712" "171568" "1716" "1717" "1718" "1719"
[205] "1723" "1737" "1738" "1743" "1757" "178"
[211] "1786" "1787" "1788" "1789" "1798" "18"
[217] "1806" "1807" "1841" "1854" "189" "1890"
[223] "1892" "191" "192134" "1962" "197258" "199857"
[229] "201595" "2023" "2026" "2027" "203" "204"
[235] "205" "2053" "2058" "210" "211" "212"
[241] "2131" "2132" "2134" "2135" "2137" "216"
[247] "217" "218" "2180" "2181" "2182" "2184"
[253] "219" "2194" "220" "2203" "221" "221223"
[259] "221823" "222" "2222" "2224" "223" "2235"
[265] "224" "226" "2271" "22845" "22856" "229"
[271] "22928" "22929" "22934" "22978" "230" "23057"
[277] "231" "23193" "23236" "23305" "23382" "23396"
[283] "23417" "23475" "23483" "23498" "23530" "23545"
[289] "23553" "23556" "2356" "23600" "23649" "23761"
[295] "239" "240" "242" "245972" "245973" "246"
[301] "246721" "247" "248" "249" "250" "251"
[307] "2523" "2524" "2525" "2526" "2527" "2528"
[313] "2529" "2530" "2531" "253558" "2538" "2539"
[319] "254531" "2548" "256435" "2571" "2572" "25796"
[325] "2581" "2582" "2583" "25834" "2584" "2585"
[331] "2588" "25885" "2589" "2590" "25902" "2591"
[337] "2592" "259230" "2593" "259307" "2595" "2597"
[343] "26007" "26035" "2618" "262" "26227" "26229"
[349] "26275" "26279" "2628" "26289" "2629" "26290"
[355] "26301" "2632" "26330" "2639" "2643" "2645"
[361] "2650" "2651" "2673" "2678" "2683" "2686"
[367] "2687" "270" "27010" "27034" "27087" "27089"
[373] "27090" "271" "2710" "2712" "27124" "27165"
[379] "272" "2720" "27235" "2729" "2730" "27306"
[385] "2731" "27349" "27430" "2744" "2746" "2747"
[391] "275" "2752" "276" "2762" "277" "278"
[397] "279" "2799" "28" "280" "2805" "2806"
[403] "2821" "283208" "283871" "284098" "284541" "2875"
[409] "290" "29071" "2937" "2954" "29796" "2987"
[415] "29880" "2990" "29906" "29920" "29922" "29925"
[421] "29926" "29929" "29947" "29958" "29968" "30"
[427] "3028" "3030" "3032" "3033" "3034" "3067"
[433] "3073" "3074" "3081" "30814" "30815" "30833"
[439] "30834" "3098" "3099" "31" "3101" "314"
[445] "3141" "3145" "3155" "3156" "3157" "3158"
[451] "316" "317749" "32" "3242" "3251" "326625"
[457] "3283" "3284" "3290" "3291" "3292" "3293"
[463] "3294" "3295" "33" "3340" "3373" "337876"
[469] "339221" "34" "340485" "341392" "3417" "3418"
[475] "3419" "341947" "3420" "3421" "3422" "3423"
[481] "3425" "348158" "349565" "35" "353" "36"
[487] "3612" "3613" "3614" "3615" "3620" "3628"
[493] "3631" "3632" "3633" "3636" "37" "3703"
[499] "3704" "3705" "3706" "3707" "3712" "374291"
[505] "374378" "3795" "38" "383" "384" "387787"
[511] "39" "3906" "391013" "3938" "3939" "3945"
[517] "3948" "3990" "4047" "4048" "4051" "4056"
[523] "411" "4121" "4122" "4124" "4128" "4129"
[529] "4143" "4144" "4190" "4191" "4199" "4245"
[535] "4247" "4248" "4249" "427" "4329" "435"
[541] "4351" "4357" "438" "440" "440138" "440567"
[547] "441024" "441531" "442117" "445" "4507" "4508"
[553] "4509" "4512" "4513" "4514" "4519" "4522"
[559] "4524" "4535" "4536" "4537" "4538" "4539"
[565] "4540" "4541" "4548" "4594" "4597" "4598"
[571] "4669" "4694" "4695" "4696" "4697" "4698"
[577] "47" "4700" "4701" "4702" "4704" "4705"
[583] "4706" "4707" "4708" "4709" "471" "4710"
[589] "4711" "4712" "4713" "4714" "4715" "4716"
[595] "4717" "4718" "4719" "4720" "4722" "4723"
[601] "4724" "4725" "4726" "4728" "4729" "4731"
[607] "48" "4830" "4831" "4832" "4833" "4837"
[613] "4842" "4843" "4846" "4860" "4907" "493911"
[619] "4942" "4952" "4953" "4967" "498" "50"
[625] "5009" "501" "5033" "5048" "50484" "50487"
[631] "5049" "5050" "5051" "5053" "506" "50614"
[637] "50617" "50700" "50814" "509" "5091" "5095"
[643] "5096" "51" "51004" "5105" "51056" "5106"
[649] "51074" "51082" "51084" "51102" "51109" "51144"
[655] "51166" "51179" "51181" "51196" "51227" "51251"
[661] "51268" "513" "5130" "51301" "51380" "51382"
[667] "514" "51477" "51478" "515" "51540" "516"
[673] "5160" "51601" "51604" "51606" "5161" "5162"
[679] "5167" "5169" "517" "51703" "51727" "51728"
[685] "51733" "51763" "518" "51805" "51809" "5198"
[691] "521" "5211" "5213" "5214" "522" "5223"
[697] "5224" "5226" "523" "5230" "5232" "5236"
[703] "525" "526" "527" "5277" "5279" "528"
[709] "5281" "5283" "5286" "5287" "5288" "5289"
[715] "529" "5297" "5298" "5313" "5315" "5319"
[721] "5320" "5321" "5322" "533" "5330" "5331"
[727] "5332" "5333" "5335" "53354" "5336" "5337"
[733] "5338" "534" "535" "53630" "537" "5372"
[739] "5373" "539" "53947" "5406" "5407" "5408"
[745] "5409" "54107" "54187" "5422" "5424" "5425"
[751] "5426" "5427" "5428" "5430" "5431" "5432"
[757] "5433" "5434" "54344" "5435" "5436" "54363"
[763] "5437" "5438" "5439" "5440" "5441" "5444"
[769] "5445" "5446" "54480" "54490" "54575" "54576"
[775] "54577" "54578" "54579" "54600" "54657" "54658"
[781] "54659" "54675" "5471" "54802" "548596" "548644"
[787] "549" "54947" "54963" "54965" "5498" "54988"
[793] "54995" "55163" "55191" "55224" "55229" "55256"
[799] "55276" "55300" "55301" "55304" "55312" "55361"
[805] "5538" "55454" "55500" "55512" "55568" "5557"
[811] "5558" "55627" "55650" "55703" "55750" "55753"
[817] "55790" "55808" "55821" "55902" "55907" "56052"
[823] "5625" "56267" "5631" "5634" "56474" "56623"
[829] "56624" "56655" "56848" "56894" "56895" "56898"
[835] "56901" "56913" "56922" "56953" "56994" "570"
[841] "57016" "57026" "57134" "5723" "5730" "5740"
[847] "5742" "5743" "57452" "574537" "57678" "57804"
[853] "57818" "57834" "5805" "5831" "5832" "5833"
[859] "58510" "5859" "586" "5860" "587" "593"
[865] "594" "5980" "60490" "60495" "6120" "6184"
[871] "6185" "622" "6240" "6241" "6296" "6303"
[877] "6307" "6309" "6342" "635" "6389" "6390"
[883] "6391" "63917" "6392" "64087" "64131" "64132"
[889] "64409" "64425" "6448" "64579" "64600" "646625"
[895] "6470" "6472" "6476" "64768" "6480" "64802"
[901] "64816" "6482" "6483" "6484" "6487" "6489"
[907] "64902" "65220" "65263" "654364" "6609" "661"
[913] "6610" "6611" "6652" "6675" "6677" "669"
[919] "6697" "6713" "6718" "6723" "683" "686"
[925] "6888" "6898" "6916" "6999" "7054" "7083"
[931] "7084" "7086" "7108" "7166" "7167" "7173"
[937] "7263" "7264" "729020" "7298" "7299" "7306"
[943] "7357" "7358" "7360" "7363" "7364" "7365"
[949] "7366" "7367" "7368" "7371" "7372" "7378"
[955] "7381" "7384" "7385" "7386" "7388" "7389"
[961] "7390" "7498" "7841" "790" "79053" "79087"
[967] "7915" "79178" "7923" "79369" "7941" "79586"
[973] "79611" "79623" "79646" "79695" "79717" "79796"
[979] "79799" "79814" "79868" "79888" "7991" "80025"
[985] "80055" "80142" "80146" "80201" "80270" "80308"
[991] "80339" "80347" "8050" "81490" "81579" "81616"
[997] "81849" "81888" "8277" "8309" "8310" "83440"
[1003] "83549" "8372" "8382" "8394" "8395" "8398"
[1009] "8399" "84002" "84076" "84172" "84245" "84265"
[1015] "84274" "84284" "84532" "84618" "84620" "84647"
[1021] "84649" "84693" "847" "84701" "84706" "84720"
[1027] "84735" "84803" "84812" "84890" "84920" "84992"
[1033] "8509" "8513" "8525" "8526" "8527" "8529"
[1039] "85365" "8540" "85465" "8560" "8564" "8566"
[1045] "8608" "8611" "8612" "8613" "8630" "8639"
[1051] "8659" "8681" "8692" "8693" "8694" "8702"
[1057] "8703" "8704" "8705" "8706" "8707" "8708"
[1063] "873" "8733" "874" "875" "8760" "8789"
[1069] "8790" "8801" "8802" "8803" "8813" "8818"
[1075] "8821" "883" "8833" "8854" "8867" "8869"
[1081] "8871" "8877" "8879" "8942" "8972" "8974"
[1087] "89869" "8992" "9" "90423" "9060" "9061"
[1093] "9091" "9114" "91373" "9162" "91734" "9197"
[1099] "9245" "92483" "9249" "9296" "93034" "93183"
[1105] "9331" "9348" "9374" "9377" "9380" "9388"
[1111] "94005" "9453" "9468" "9487" "9488" "9489"
[1117] "95" "9514" "9517" "952" "9533" "9536"
[1123] "9550" "9551" "9563" "9588" "9615" "978"
[1129] "9791" "9942" "9945"
$`hsa00230 Purine metabolism`
[1] "100" "10201" "10606" "10621" "10622" "10623" "107" "10714"
[9] "108" "10846" "109" "111" "11128" "11164" "112" "113"
[17] "114" "115" "122481" "122622" "124583" "132" "158" "159"
[25] "1633" "171568" "1716" "196883" "203" "204" "205" "221823"
[33] "2272" "22978" "23649" "246721" "25885" "2618" "26289" "270"
[41] "271" "27115" "272" "2766" "2977" "2982" "2983" "2984"
[49] "2986" "2987" "29922" "3000" "30833" "30834" "318" "3251"
[57] "353" "3614" "3615" "3704" "377841" "471" "4830" "4831"
[65] "4832" "4833" "4860" "4881" "4882" "4907" "50484" "50940"
[73] "51082" "51251" "51292" "5136" "5137" "5138" "5139" "5140"
[81] "5141" "5142" "5143" "5144" "5145" "5146" "5147" "5148"
[89] "5149" "5150" "5151" "5152" "5153" "5158" "5167" "5169"
[97] "51728" "5198" "5236" "5313" "5315" "53343" "54107" "5422"
[105] "5424" "5425" "5426" "5427" "5430" "5431" "5432" "5433"
[113] "5434" "5435" "5436" "5437" "5438" "5439" "5440" "5441"
[121] "5471" "548644" "55276" "5557" "5558" "55703" "55811" "55821"
[129] "5631" "5634" "56655" "56953" "56985" "57804" "58497" "6240"
[137] "6241" "64425" "646625" "654364" "661" "7498" "8382" "84172"
[145] "84265" "84284" "84618" "8622" "8654" "87178" "8833" "9060"
[153] "9061" "93034" "953" "9533" "954" "955" "956" "957"
[161] "9583" "9615"
$`hsa05340 Primary immunodeficiency`
[1] "100" "115650" "23495" "29760" "29851" "326" "3543" "3561"
[9] "3575" "3718" "3932" "4261" "57379" "5788" "5896" "5897"
[17] "5993" "5994" "64421" "6890" "6891" "695" "7374" "7535"
[25] "8517" "8625" "915" "916" "920" "925" "926" "930"
[33] "958" "959" "973" keggres <- gage(foldchanges, gsets=kegg.sets.hs)
head(keggres$less) p.geomean stat.mean
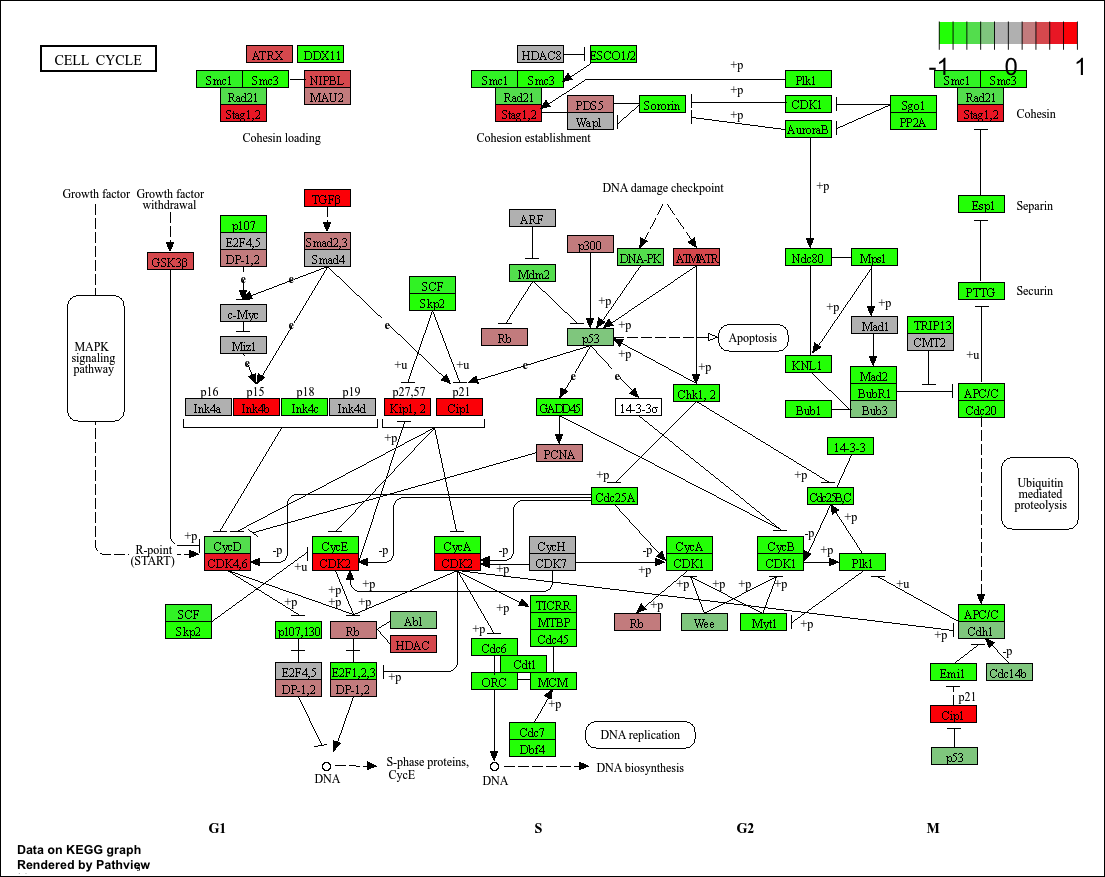
hsa04110 Cell cycle 8.995727e-06 -4.378644
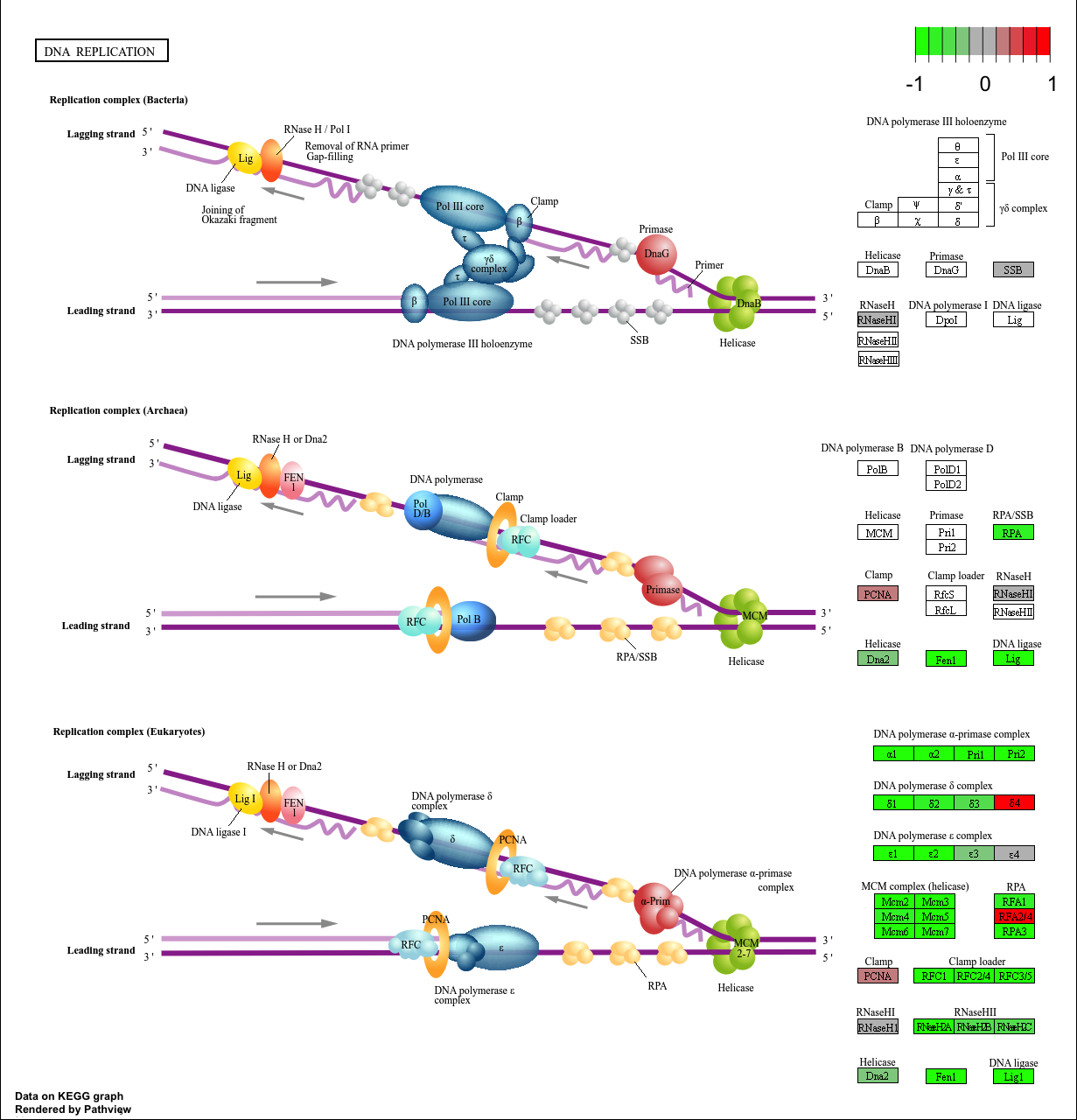
hsa03030 DNA replication 9.424076e-05 -3.951803
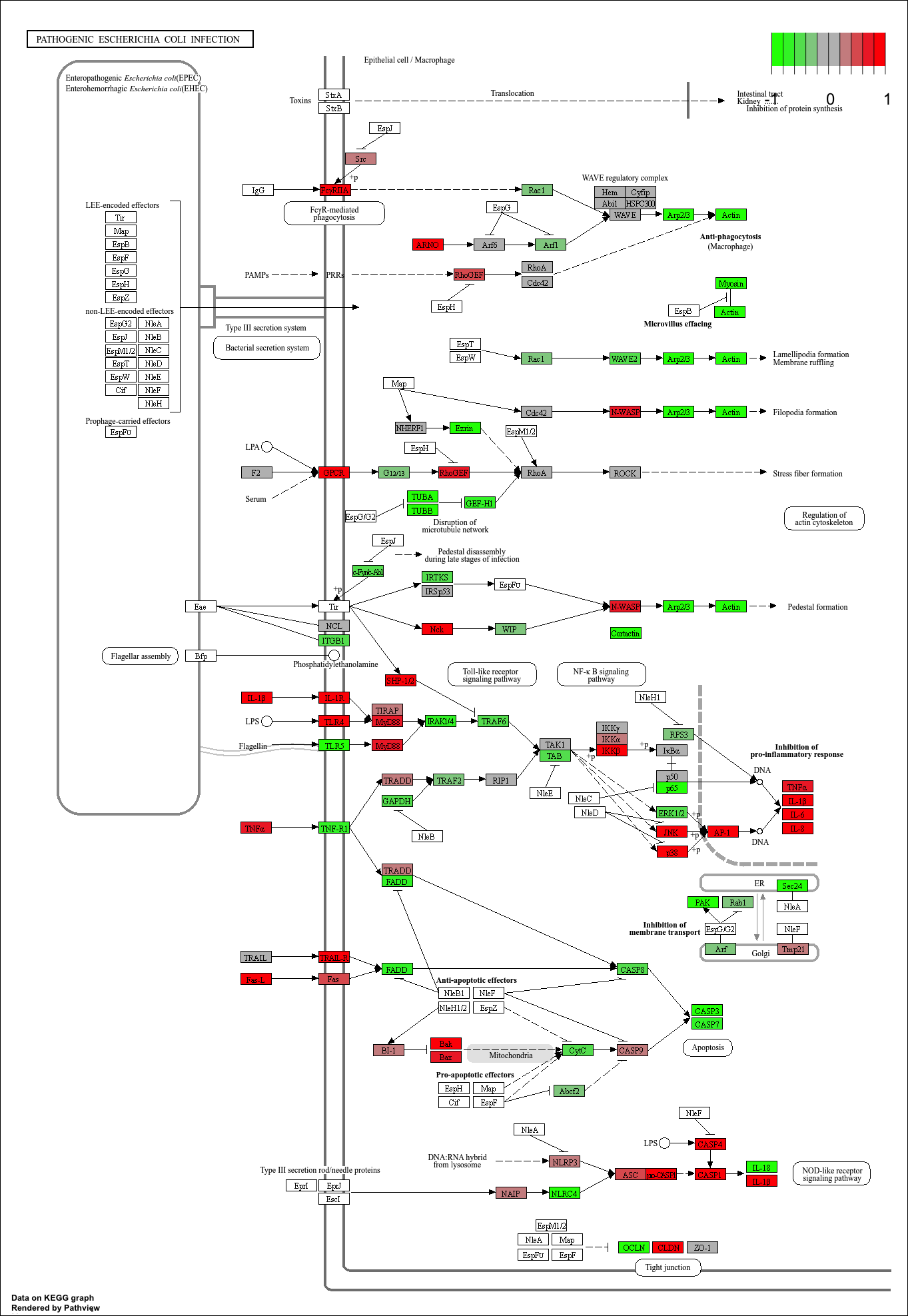
hsa05130 Pathogenic Escherichia coli infection 1.405864e-04 -3.765330
hsa03013 RNA transport 1.375901e-03 -3.028500
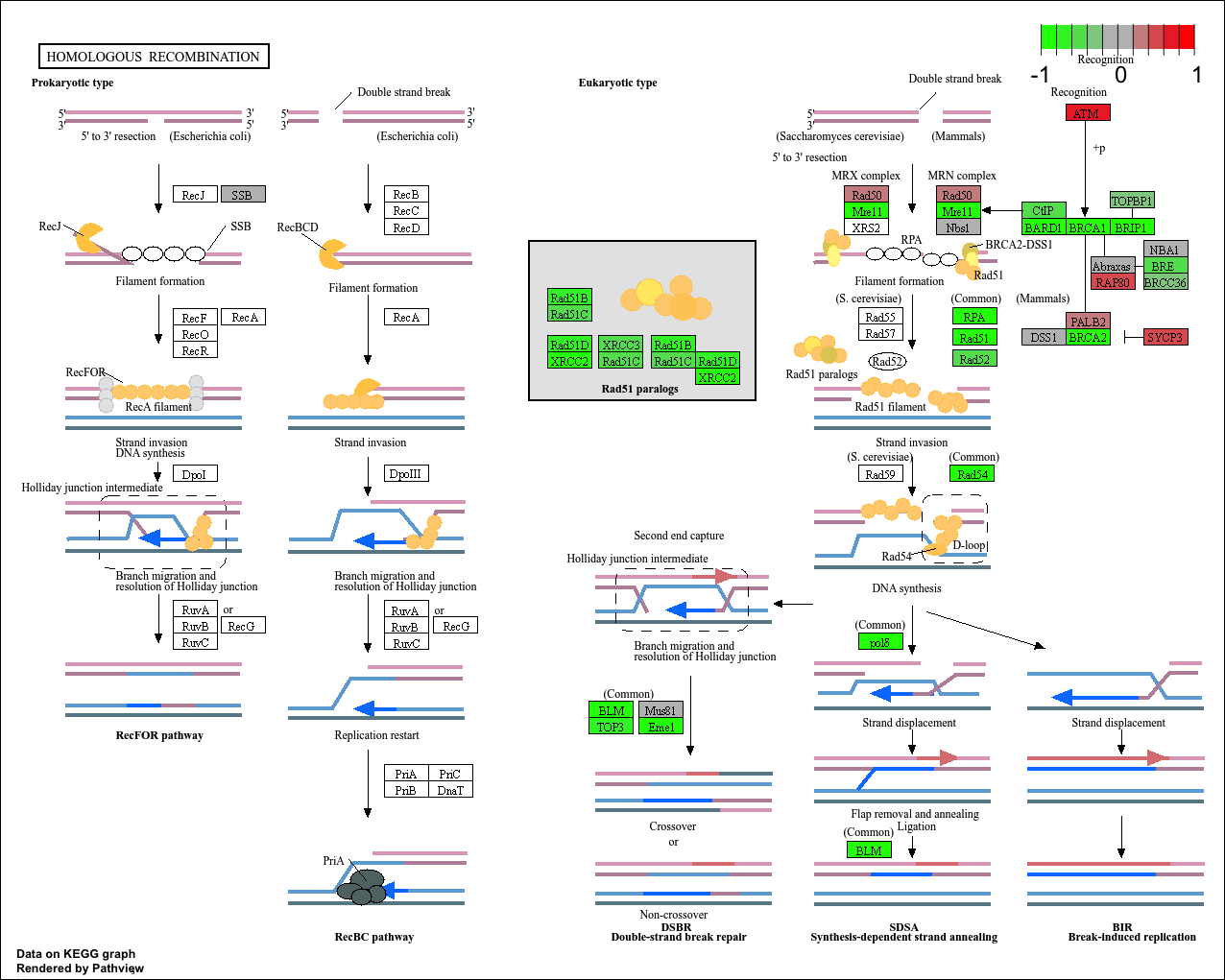
hsa03440 Homologous recombination 3.066756e-03 -2.852899
hsa04114 Oocyte meiosis 3.784520e-03 -2.698128
p.val q.val
hsa04110 Cell cycle 8.995727e-06 0.001889103
hsa03030 DNA replication 9.424076e-05 0.009841047
hsa05130 Pathogenic Escherichia coli infection 1.405864e-04 0.009841047
hsa03013 RNA transport 1.375901e-03 0.072234819
hsa03440 Homologous recombination 3.066756e-03 0.128803765
hsa04114 Oocyte meiosis 3.784520e-03 0.132458191
set.size exp1
hsa04110 Cell cycle 121 8.995727e-06
hsa03030 DNA replication 36 9.424076e-05
hsa05130 Pathogenic Escherichia coli infection 53 1.405864e-04
hsa03013 RNA transport 144 1.375901e-03
hsa03440 Homologous recombination 28 3.066756e-03
hsa04114 Oocyte meiosis 102 3.784520e-03KEGG
keggrespathways <- rownames(keggres$less)[1:5]
keggresids <- substr(keggrespathways, start=1, stop=8)
keggresids[1] "hsa04110" "hsa03030" "hsa05130" "hsa03013" "hsa03440"pathview(gene.data=foldchanges, pathway.id=keggresids, species="hsa")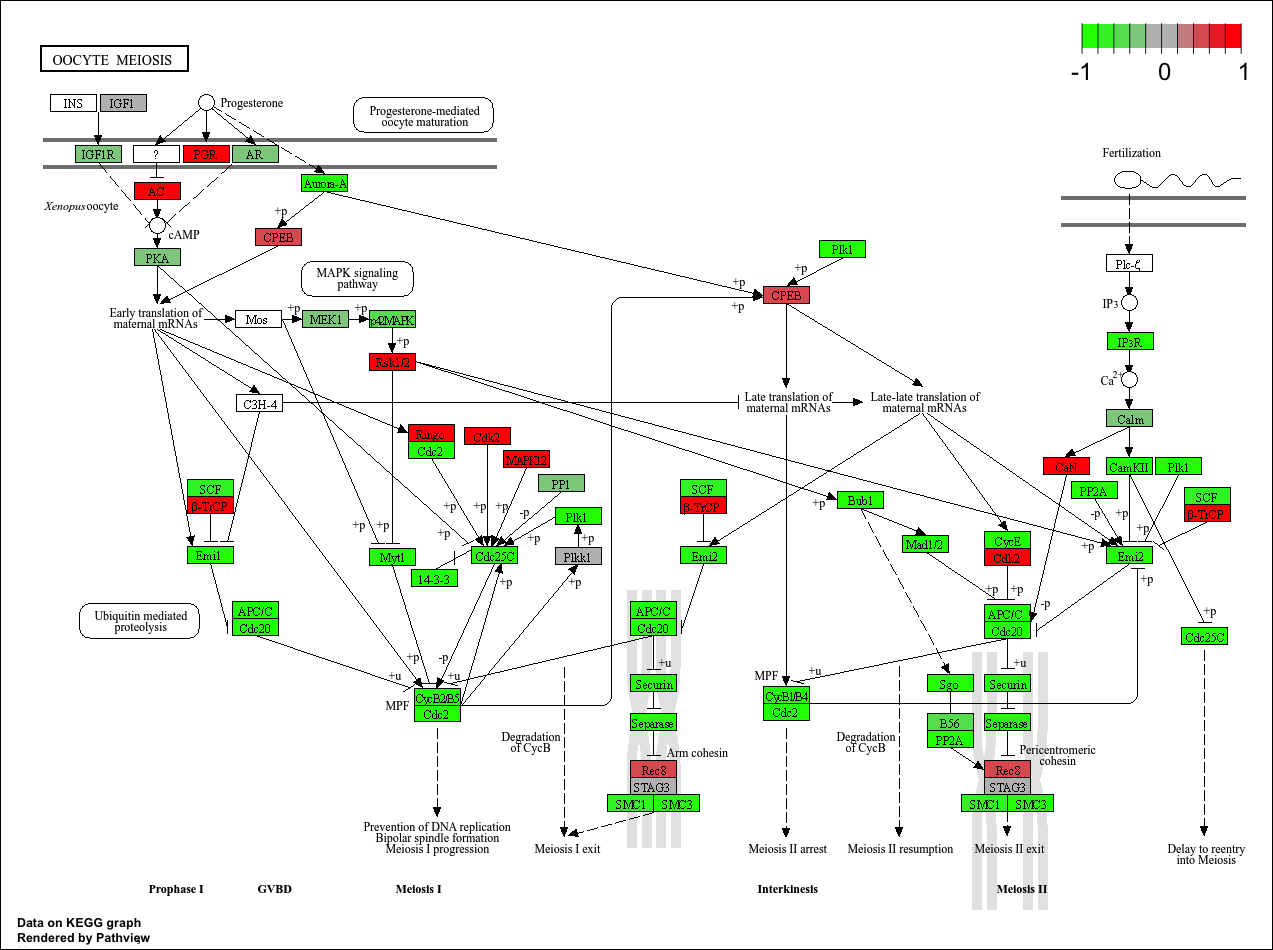
GO
data(go.sets.hs)
data(go.subs.hs)
gobpsets = go.sets.hs[go.subs.hs$BP]
gobpres = gage(foldchanges, gsets=gobpsets)
lapply(gobpres, head)$greater
p.geomean stat.mean p.val
GO:0007156 homophilic cell adhesion 8.519724e-05 3.824205 8.519724e-05
GO:0002009 morphogenesis of an epithelium 1.396681e-04 3.653886 1.396681e-04
GO:0048729 tissue morphogenesis 1.432451e-04 3.643242 1.432451e-04
GO:0007610 behavior 1.925222e-04 3.565432 1.925222e-04
GO:0060562 epithelial tube morphogenesis 5.932837e-04 3.261376 5.932837e-04
GO:0035295 tube development 5.953254e-04 3.253665 5.953254e-04
q.val set.size exp1
GO:0007156 homophilic cell adhesion 0.1951953 113 8.519724e-05
GO:0002009 morphogenesis of an epithelium 0.1951953 339 1.396681e-04
GO:0048729 tissue morphogenesis 0.1951953 424 1.432451e-04
GO:0007610 behavior 0.1967577 426 1.925222e-04
GO:0060562 epithelial tube morphogenesis 0.3565320 257 5.932837e-04
GO:0035295 tube development 0.3565320 391 5.953254e-04
$less
p.geomean stat.mean p.val
GO:0048285 organelle fission 1.536227e-15 -8.063910 1.536227e-15
GO:0000280 nuclear division 4.286961e-15 -7.939217 4.286961e-15
GO:0007067 mitosis 4.286961e-15 -7.939217 4.286961e-15
GO:0000087 M phase of mitotic cell cycle 1.169934e-14 -7.797496 1.169934e-14
GO:0007059 chromosome segregation 2.028624e-11 -6.878340 2.028624e-11
GO:0000236 mitotic prometaphase 1.729553e-10 -6.695966 1.729553e-10
q.val set.size exp1
GO:0048285 organelle fission 5.841698e-12 376 1.536227e-15
GO:0000280 nuclear division 5.841698e-12 352 4.286961e-15
GO:0007067 mitosis 5.841698e-12 352 4.286961e-15
GO:0000087 M phase of mitotic cell cycle 1.195672e-11 362 1.169934e-14
GO:0007059 chromosome segregation 1.658603e-08 142 2.028624e-11
GO:0000236 mitotic prometaphase 1.178402e-07 84 1.729553e-10
$stats
stat.mean exp1
GO:0007156 homophilic cell adhesion 3.824205 3.824205
GO:0002009 morphogenesis of an epithelium 3.653886 3.653886
GO:0048729 tissue morphogenesis 3.643242 3.643242
GO:0007610 behavior 3.565432 3.565432
GO:0060562 epithelial tube morphogenesis 3.261376 3.261376
GO:0035295 tube development 3.253665 3.253665