counts <- read.csv("airway_scaledcounts.csv", row.names=1)
metadata <- read.csv("airway_metadata.csv")Class 13 Transcriptomics and the analysis of RNA-Seq data
Background
Today we will perform an RNASeq analysis on the effects of dexamethasone (hereafter “dex”), a common steroid, on airway smooth muscle (ASM) cell lines.
Data Import
We need two things for this analysis:
- countData: a table with genes as rows and samples/experiments as columns,
- colData: metadata about the columns (i.e. samples) in the main countData object.
Let’s have a wee peak at these two objects:
metadata id dex celltype geo_id
1 SRR1039508 control N61311 GSM1275862
2 SRR1039509 treated N61311 GSM1275863
3 SRR1039512 control N052611 GSM1275866
4 SRR1039513 treated N052611 GSM1275867
5 SRR1039516 control N080611 GSM1275870
6 SRR1039517 treated N080611 GSM1275871
7 SRR1039520 control N061011 GSM1275874
8 SRR1039521 treated N061011 GSM1275875head(counts) SRR1039508 SRR1039509 SRR1039512 SRR1039513 SRR1039516
ENSG00000000003 723 486 904 445 1170
ENSG00000000005 0 0 0 0 0
ENSG00000000419 467 523 616 371 582
ENSG00000000457 347 258 364 237 318
ENSG00000000460 96 81 73 66 118
ENSG00000000938 0 0 1 0 2
SRR1039517 SRR1039520 SRR1039521
ENSG00000000003 1097 806 604
ENSG00000000005 0 0 0
ENSG00000000419 781 417 509
ENSG00000000457 447 330 324
ENSG00000000460 94 102 74
ENSG00000000938 0 0 0Check on metadata counts correspondance
We need to check that the metadata matches the samples in our count data.
ncol(counts) == nrow(metadata)[1] TRUEcolnames(counts) == metadata$id[1] TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUEall( c(T,T,T,T))[1] TRUEQ1. how many genes are in this dataset?
nrow(counts)[1] 38694Q2. How many “control” samples are in this dataset?
sum(metadata$dex == "control")[1] 4Analysis Plan…
We have 4 replicates per condition (“control” and “treated”). We want to compare the control vs the treated to see which genes expression levels change when we have the drug present.
We will go row by row (gene by gene) and see if the average value in control columns is different than the average value in treated columns
Step 1. Find which columns in
countscorrespond to “control” samples.Step 2. Extract/select these columns
Step 3. Calculate an average value for each gene (i.e. each row).
# The indices (i.e positions) that are "control"
control.inds <- metadata$dex == "control"# Extract/select these "control" columns from counts
control.counts <- counts[,control.inds]# Calculate the mean for each gene (i.e row)
control.mean <- rowMeans(control.counts)Q. Do the same for “treated” samples - find the mean count value per gene
treated.inds <- metadata$dex == "treated"
treated.counts <- counts[,treated.inds]
treated.mean <- rowMeans(treated.counts)Let’s put these two mean values into a new data.frame meancounts for easy book-keeping and plotting.
meancounts <- data.frame(control.mean, treated.mean)
head(meancounts) control.mean treated.mean
ENSG00000000003 900.75 658.00
ENSG00000000005 0.00 0.00
ENSG00000000419 520.50 546.00
ENSG00000000457 339.75 316.50
ENSG00000000460 97.25 78.75
ENSG00000000938 0.75 0.00Q. Make a ggplot of average counts of control vs treated.
library(ggplot2)
ggplot(meancounts, aes(control.mean, treated.mean)) + geom_point(alpha = 0.3) + scale_x_log10() + scale_y_log10()Warning in scale_x_log10(): log-10 transformation introduced infinite values.Warning in scale_y_log10(): log-10 transformation introduced infinite values.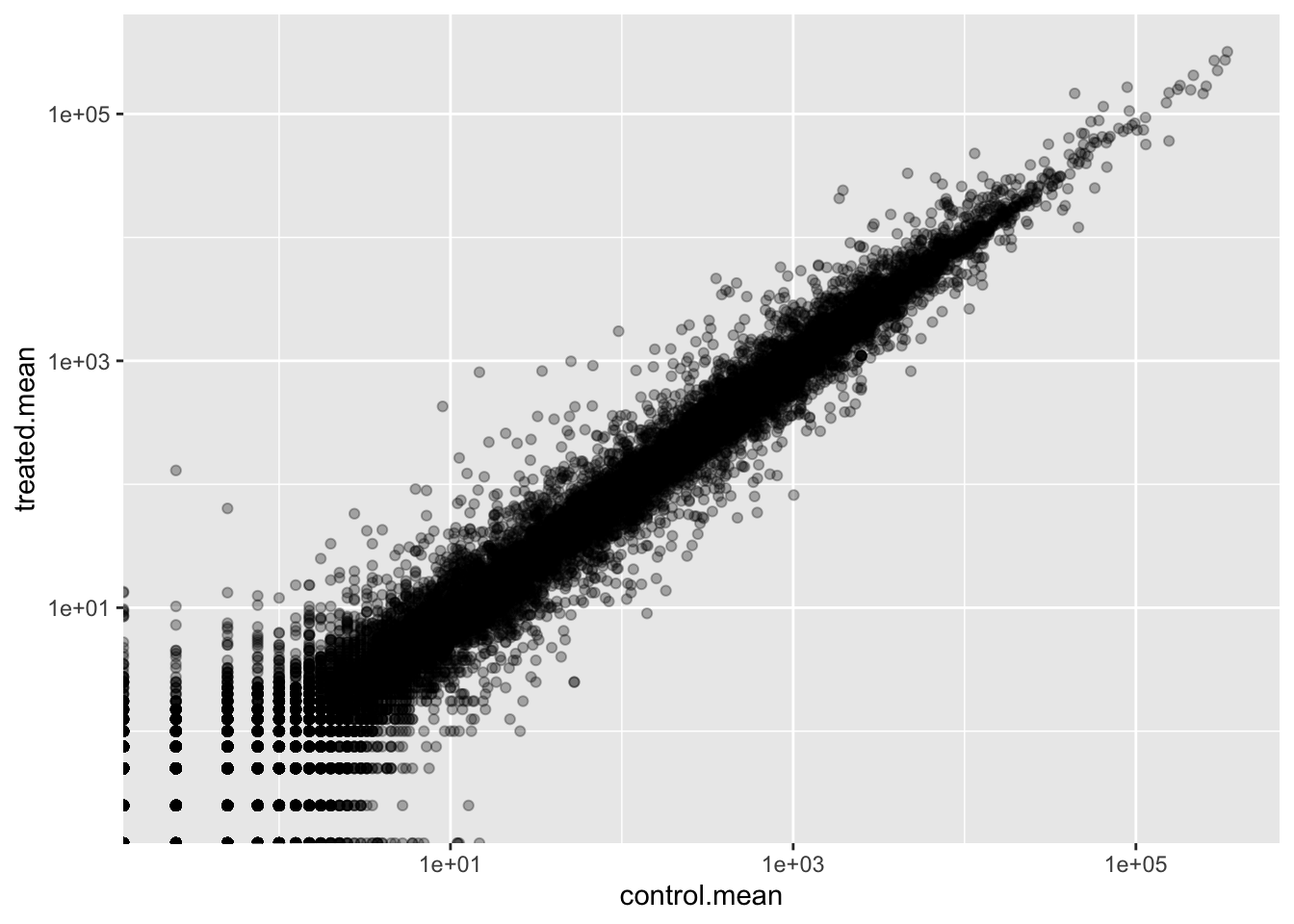
Log2 units and fold change
If we consider “treated”/ “control” counts we will get a number that tells us the change.
# No change
log2(20/20)[1] 0# A doubling in the treated vs control
log2(40/20)[1] 1log2(10/40)[1] -2Q. Add a new column
log2fcfor log2 fold change of treated/control to ourmeancountsobject.
meancounts$log2fc <-
log2(meancounts$treated.mean/meancounts$control.mean)
head(meancounts) control.mean treated.mean log2fc
ENSG00000000003 900.75 658.00 -0.45303916
ENSG00000000005 0.00 0.00 NaN
ENSG00000000419 520.50 546.00 0.06900279
ENSG00000000457 339.75 316.50 -0.10226805
ENSG00000000460 97.25 78.75 -0.30441833
ENSG00000000938 0.75 0.00 -InfRemove zero count genes
Typically we would not consider zero count genes - as we have no data about them and they should be excluded from further consideration. These lead to “funky” log2 fold change values (e.g. divide by zero errors etc.)
DESeq analysis
We are missing any measure of significance from the work we had so far. Let’s do this properly with the DESeq2 package.
library(DESeq2)The DESeq2 package, like many bioconductor packages, wants it’s input in a very specific way - a data structure setup with all the info it needs for the calculation.
dds <- DESeqDataSetFromMatrix(countData = counts, colData = metadata, design = ~dex)converting counts to integer modeWarning in DESeqDataSet(se, design = design, ignoreRank): some variables in
design formula are characters, converting to factorsThe main function in this package is called DESeq() it will run the full analysis for us on our dds input object:
dds <- DESeq(dds)estimating size factorsestimating dispersionsgene-wise dispersion estimatesmean-dispersion relationshipfinal dispersion estimatesfitting model and testingExtract our results:
res <-results(dds)
head(res)log2 fold change (MLE): dex treated vs control
Wald test p-value: dex treated vs control
DataFrame with 6 rows and 6 columns
baseMean log2FoldChange lfcSE stat pvalue
<numeric> <numeric> <numeric> <numeric> <numeric>
ENSG00000000003 747.194195 -0.3507030 0.168246 -2.084470 0.0371175
ENSG00000000005 0.000000 NA NA NA NA
ENSG00000000419 520.134160 0.2061078 0.101059 2.039475 0.0414026
ENSG00000000457 322.664844 0.0245269 0.145145 0.168982 0.8658106
ENSG00000000460 87.682625 -0.1471420 0.257007 -0.572521 0.5669691
ENSG00000000938 0.319167 -1.7322890 3.493601 -0.495846 0.6200029
padj
<numeric>
ENSG00000000003 0.163035
ENSG00000000005 NA
ENSG00000000419 0.176032
ENSG00000000457 0.961694
ENSG00000000460 0.815849
ENSG00000000938 NAVolcano plot
A useful summary figure of our results is often called a volcano pot. It is basically a plot of log2 fold change values vs Adjusted p-values.
Q. use ggplot to make a first version “volcano plot” of
log2FoldChangevspadj
ggplot(res, aes(log2FoldChange, padj)) + geom_point()Warning: Removed 23549 rows containing missing values or values outside the scale range
(`geom_point()`).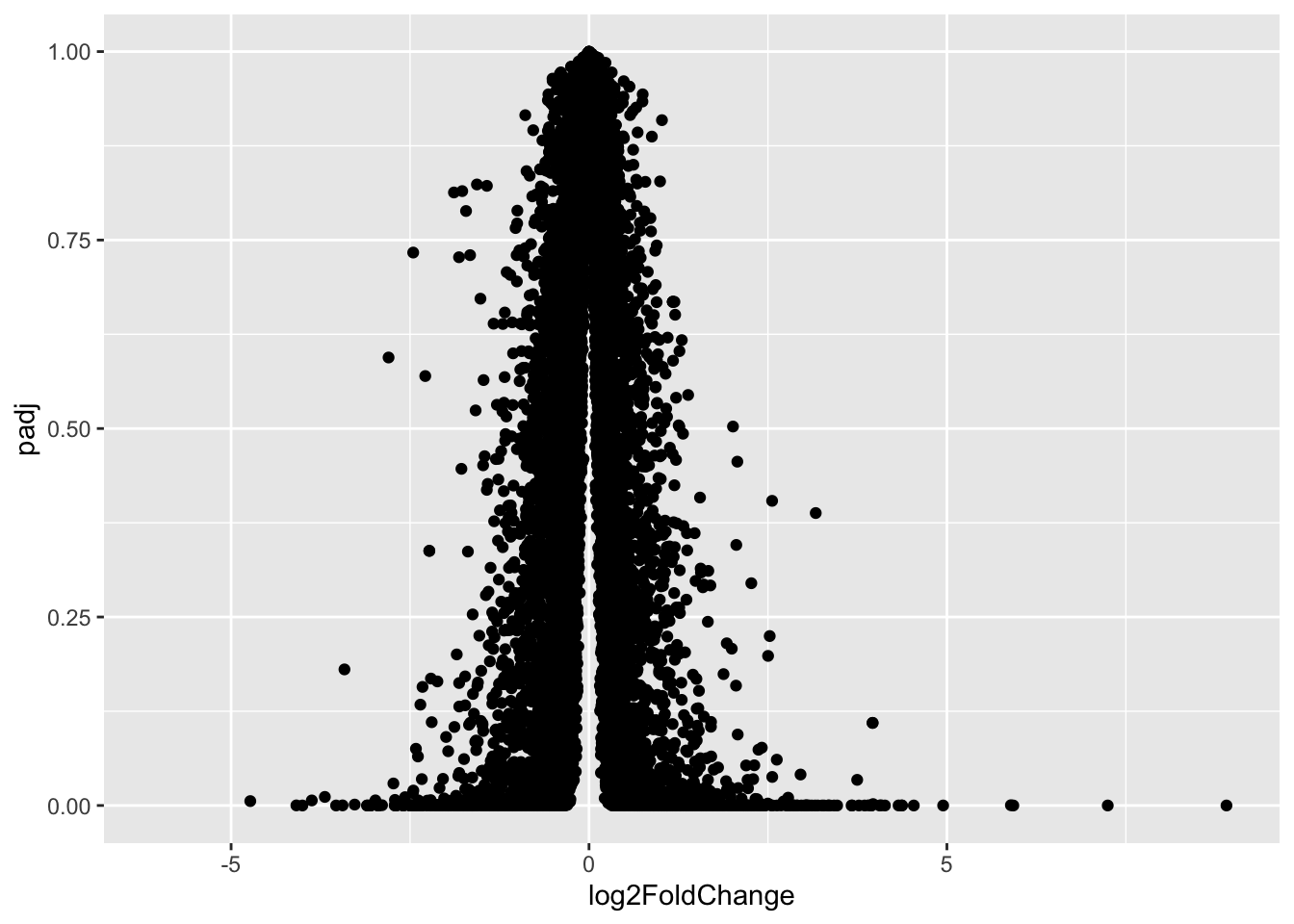
This is not very useful because the y-axis (p-value) is not really helpful - we want to focus on low p-values
ggplot(res, aes(log2FoldChange, log(padj))) + geom_point()Warning: Removed 23549 rows containing missing values or values outside the scale range
(`geom_point()`).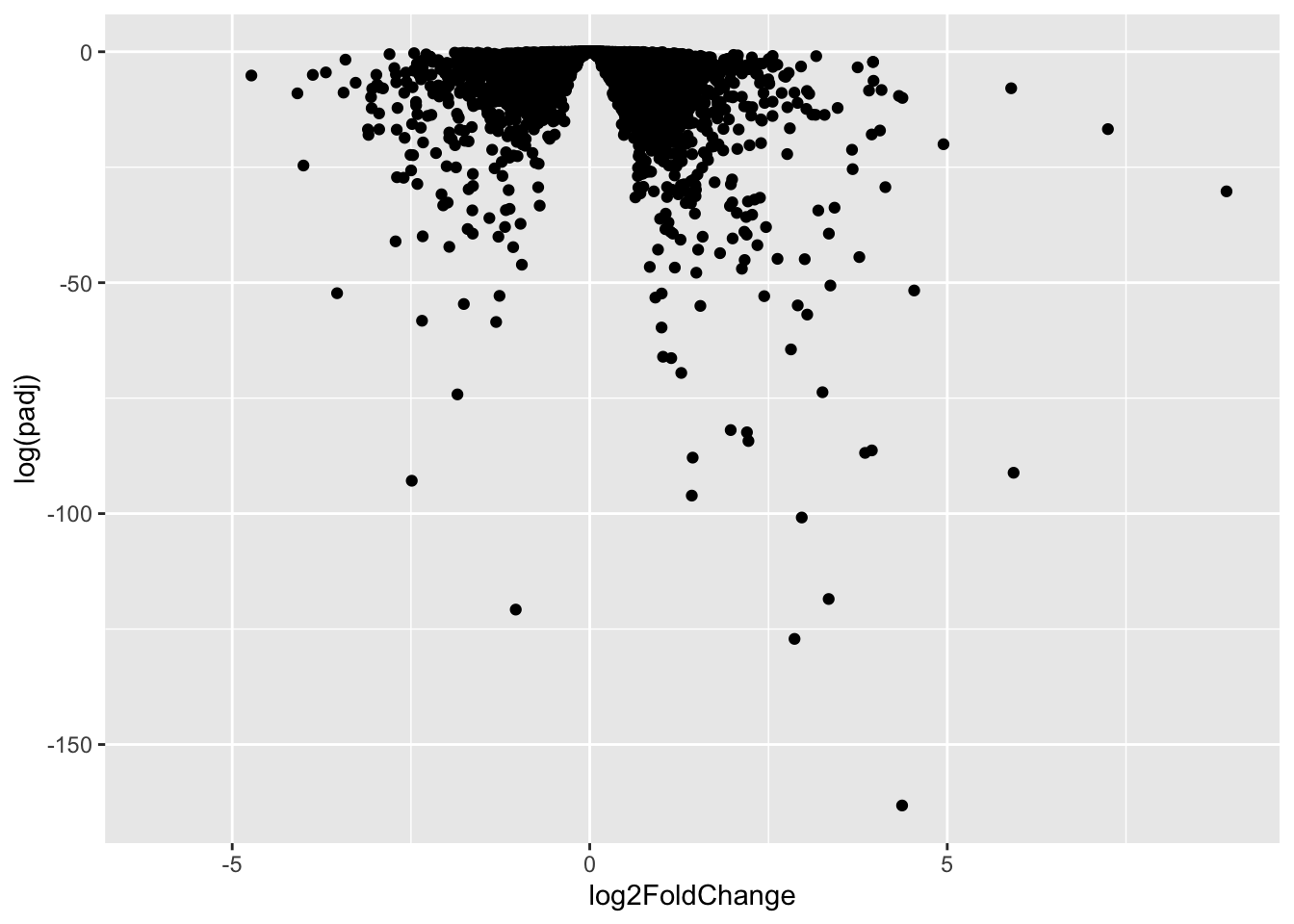
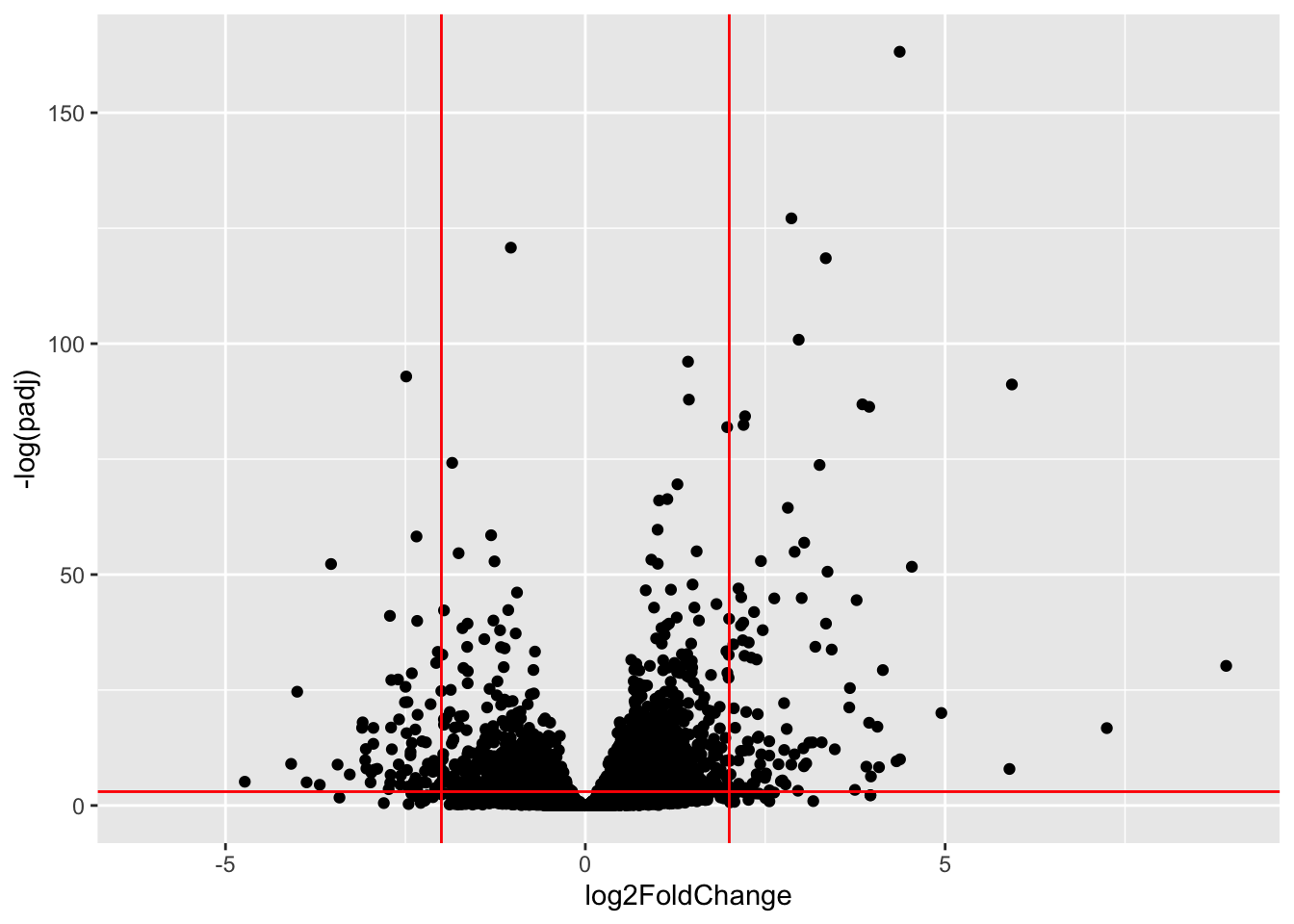
ggplot(res, aes(log2FoldChange, -log(padj))) + geom_point() + geom_vline(xintercept = c(-2,+2), col = "red") + geom_hline(yintercept = -log(0.05), col = "red")Warning: Removed 23549 rows containing missing values or values outside the scale range
(`geom_point()`).